Bacteriophage Discovery & Characterization
Discovery and characterization of a novel soil bacteriophage through authentic research.
Program: SEA-PHAGES Undergraduate Research Program
Course Context: Research Laboratory (Two-Person Team)
Role: Soil sampling, phage isolation & purification, titer analysis, DNA extraction, data submission
Tools & Techniques: Plaque assays, serial dilution, PCR, spectrophotometry, TEM, aseptic technique
Output: Public database submission, scientific poster & symposium presentation
Overview
This is a research laboratory project that was undertaken as part of the SEA-PHAGES (Science Education Alliance-Phage Hunters Advancing Genomics and Evolutionary Science) undergraduate research program. The objective was to isolate, purify and characterize a previously undocumented bacteriophage from natural soil samples.
Working as part of a two-person research team, I completed the full experimental pipeline, including environmental sampling and microbial isolation, genomic preparation, and submission of publicly available data. The resulting bacteriophage was effectively described and deposited in the Actinobacteriophage Database as Madique, which is a part of a world-wide open-access research database.
In addition to microbiology, this project focused on experimental design, quantitative analysis and data validation-skills that are directly relevant to environmental engineering research, bioremediation, and microbial monitoring.
Visit The Actinobacteriophage Database entry: Madique
Objectives
Isolate a novel bacteriophage from a natural soil environment
Purify a single genetically distinct phage strain
Quantify phage concentration using plaque assays and serial dilutions
Isolate, extract and prepare high-quality phage DNA for sequencing
Characterize phage morphology using Transmission Electron Microscopy (TEM)
Disseminate findings through a scientific database and poster presentation
Methods
Soil Sampling and Documentation.
Soil samples were collected from a local site in South Florida. Environmental parameters including GPS coordinates, temperature, pH, and weather conditions were recorded to contextualize microbial diversity.
Isolation and Enrichment
Both direct and enriched isolations were performed using Mycobacterium foliorum as the bacterial host. Phage presence was confirmed by the formation of plaques indicating bacterial lysis. Plaques were purified through three successive rounds of isolation to ensure genetic purity.
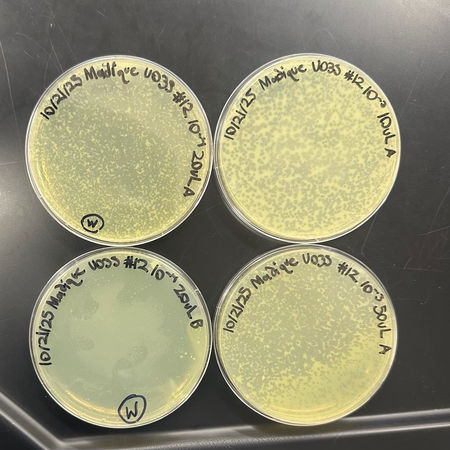
Purification and Plaque Assay.
Purified plaque thrice to be sure that only one, genetically pure phage strain was isolated. Serial dilution and top-agar plating were used each round to observe the morphology of the plaque and estimate titer.
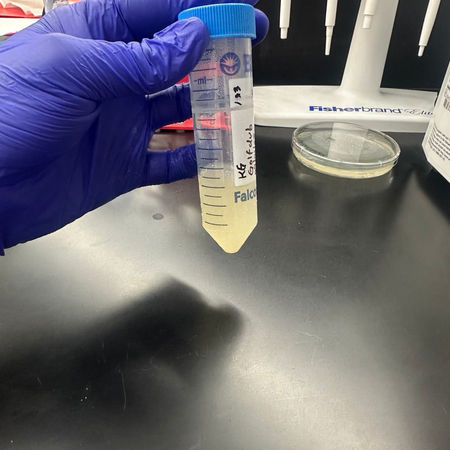
Lysate Collection and Titer Calculation.
Growth of high-titer lysate by flooding webbed plates and spot titer and full-plate titer assays to establish the concentration of the phage (plaque-forming units per milliliter).
DNA Extraction and Quantification
The DNA was extracted and quantified by the direct PCR method using the primers, DNA polymerase chain reaction (PCR) mixture, and the DNA polymerase chain reaction (PCR) product. Purified and isolated DNA through a resin-based protocol, and then spectrophotometrically quantified to verify purity and yield to prepare a sequencing sample.
Transmission Electron Microscopy (TEM)
Images of phage capsid and tail morphology analyzed using captured high-resolution images. The long flexible tail of Madique was siphovirus-like, which is typical of phages infecting Mycobacterium hosts.
Data Submission and Presentation.
Submitted final results to PhagesDB and gave a presentation in the Phage Olympics, the closing research symposium of the program.



Results
Phage Identified: Madique
Host: Mycobacterium foliorum
Stain: Uranyl acetate
Magnification ~60,000x
Titer: 6.0 x 108 PFU/mL
Morphology Summary (TEM):
Classic Podoviridae-like morphology
Icosahedral capsid: ~60 nm diameter.
Short, non-contractile tail: ~20nm.
No visible sheath.
No ruptures or artifacts

Madique exhibits the morphology of a double-stranded DNA bacteriophage with an icosahedral capsid (~60 nm) and a short, non-contractile tail (~20 nm). These features are consistent with podovirus-like actinobacteriophages isolated using Mycobacterium hosts in the SEA-PHAGES program.
Engineering Relevance
Although this project has its foundation in molecular biology, it is directly parallel to the research approaches of environmental engineering:
Sampling & Characterization: As with microbial monitoring in soil and water quality systems.
Quantification Techniques: The method of dilution and assay is a reflection of pollutant or microbial quantification.
Bioinformatics & Data Integration: PhagesDB process presents data management and open-source collaboration- required in the contemporary environmental modeling.
Applied Implications: The dynamics of bacteriophages can be used to gain an insight into microbial ecology, wastewater treatment, and bio-control of pathogens in the natural environment.
Reflection
This laboratory helped develop technical and analytical expertise that is essential in engineering research. It needed accuracy, trial and error, and evidence-based decision-making. In addition to proficiency in the laboratory, it enhanced a comprehension of the role of microbial interactions on environmental systems to bridge the gap between molecular biology and environmental design.
The isolation and characterization of Madique provided me with a physical feeling of discovery- turning an invisible piece of soil into a genetically distinct publicly documented organism. This experience reinforced my interest in the combination of biological research and environmental engineering innovation.
Project Gallery